Reinhard Windoffer and Rudolf Leube with Sonja Lehmann, Jana Schieren and Sungjung Yoon
Keratin intermediate filaments are major constituents of the epithelial cytoskeleton. Their organizing function of cellular space is important for diverse epithelial functions requiring a high degree of keratin network plasticity. We have recently proposed that a continuous cycle of keratin filament assembly and disassembly (Figure 1) affords rapid and precise adaptation to the different functional requirements and situations. The concept is based on experiments observing fluorescently-tagged keratin polypeptides allowing us to quantitatively measure keratin network properties, speed of keratin movement and local turnover. We further demonstrated that differences in keratin cycling take place after seeding of cells and upon growth factor addition, migration induction, keratin phosphorylation, and downregulation of cytoskeletal cross linkers as well as in response to altered substrate stiffness.
Our current goal is obtain a molecular understanding of keratin network dynamics by examining candidate modulators, by studying the effect of molecular motors on keratin motility and by genetic screening. Consequences of keratin network modulation on epithelial function, i.e. cell migration, cellular viscoelasticity and stiffness, proliferation and stress response are being investigated. Our long-term goal is to understand molecular mechanisms of keratin network plasticity in the context of epithelial pathophysiology.
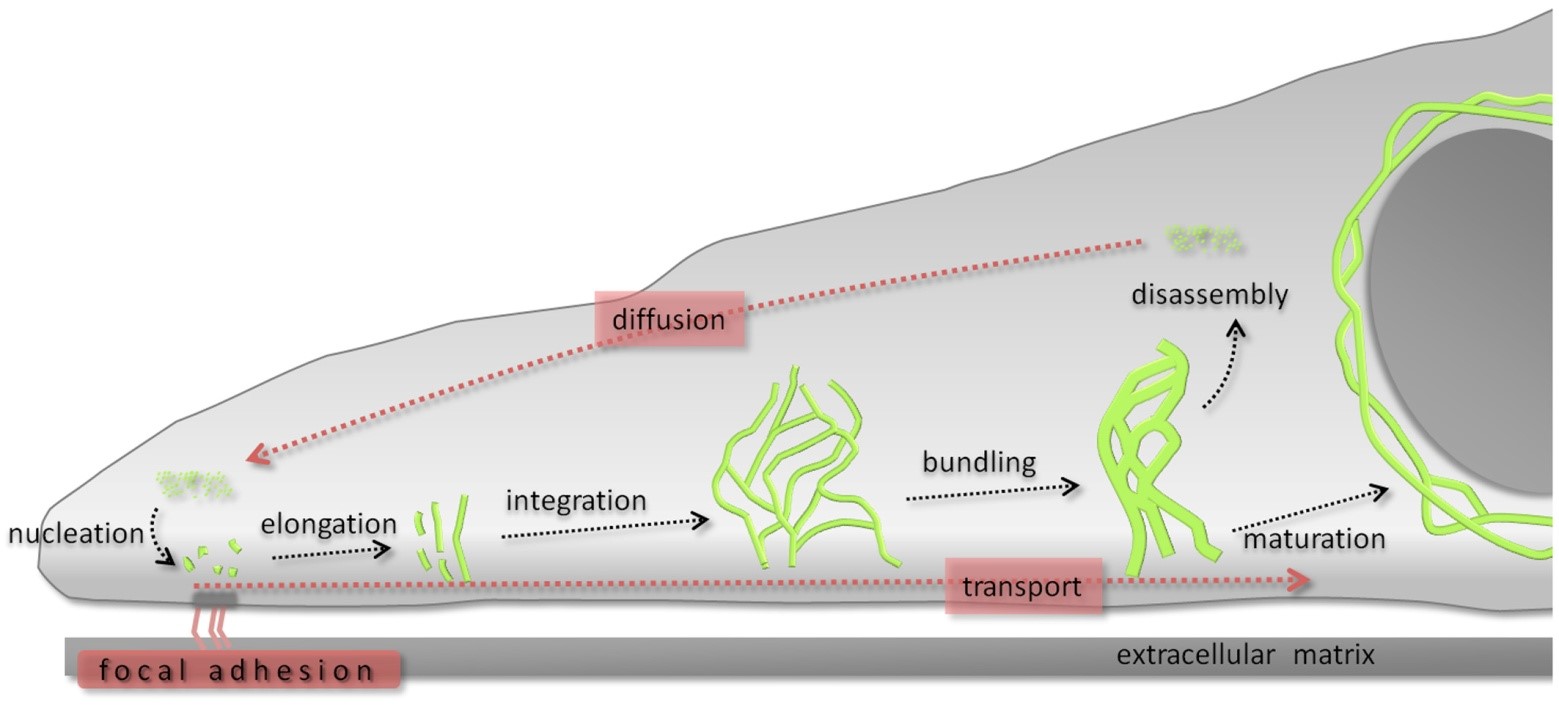
Figure 1. The keratin cycle. Soluble keratin oligomers assemble into particles in the cell periphery in proximity to focal adhesion sites (nucleation). These particles grow (elongation) and move toward the cell center in an actin-dependent process (transport). Subsequently, elongated keratin filament particles are incorporated into the peripheral keratin filament network (integration). Filament bundling occurs during further centripetal translocation toward the nucleus (transport). Soluble oligomers dissociate (disassembly), diffuse throughout the cytoplasm (diffusion), and are reutilized for another cycle of keratin filament formation in the cell periphery. Alternatively, bundled filaments are stabilized (maturation), forming, e.g., the stable perinuclear cage. (The figure is taken from Windoffer et al, 2011, J Cell Biol 194, 669.)
Cytoplasmic protein aggregates are hallmarks of many diseases, which affect various organs and lead to diverse pathologies such as neurodegeneration, myopathy or epidermal blistering. The protein composition of these aggregates varies considerably but intermediate filament polypeptides have been shown to be involved in most instances. The view that the protein aggregates are simply static cellular debris is increasingly questioned and various studies support the view that they are quite dynamic. This has kindled interest in understanding and modulating the dynamic processes of protein aggregate formation in order to interfere with disease initiation and progression for the benefit of affected patients.
Our focus is on epidermolysis bullosa simplex (EBS), which is a prototypic protein aggregate-forming disease affecting the epidermis. The autosomal-dominant disease is caused by mutations in keratin 5 and keratin 14. It manifests with severe blistering in response to mild mechanical trauma because of cytolysis in the basal cell layer of the epidermis. At the subcellular level, EBS is characterized by disruption of the keratin filament network and occurrence of keratin 5- and 14-positive amorphous cytoplasmic aggregates.
Our research is based on the observation that EBS-mutant keratins also lead to the formation of large cytoplasmic aggregates in cultured cells with unique signet features and consequences on cellular function (Figure 1). We aim to elucidate the mechanisms leading to aggregate formation and the associated pathology.
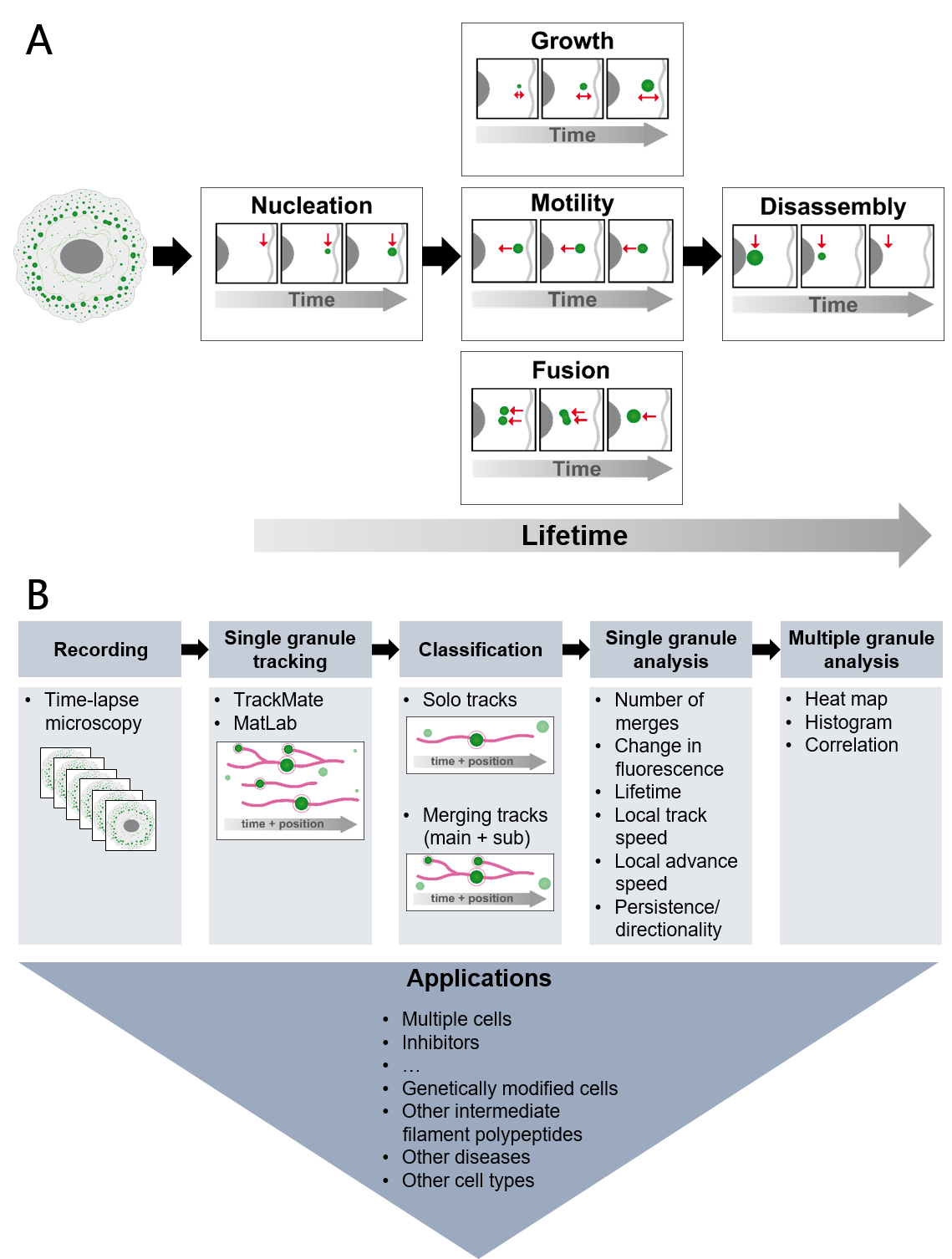
Figure 1 The schemes present properties of mutant keratin granules that can be deduced from high resolution time-lapse imaging of epithelial cells producing fluorescence-tagged keratins (A) and the workflow to deduce the properties together with possible applications (B). (The figure is taken from Lehmann et al, 2021, Sci Rep 11:2379)